Fitting#
Fitting data to a model is a very common task in physics that is non-trivial. Here we’ll go through a few ways to fit data and some common pitfalls.
There are a number of python fitting modules out there and we’ll using iminuit for this class as it neatly allows the consideration of uncertainties.
%pip install iminuit
%pip install numba_stats
from matplotlib import pyplot as plt
import numpy as np
from iminuit import Minuit
from iminuit.cost import LeastSquares
import iminuit
import seaborn as sns; sns.set()
import time
def pdf_demo(xlo, xhi, **kwargs):
x = np.linspace(xlo, xhi, 200)
cmap = sns.color_palette().as_hex()
for i, name in enumerate(kwargs):
for j, arglist in enumerate(kwargs[name].split(';')):
args = eval('dict(' + arglist + ')')
y = eval('scipy.stats.' + name)(**args).pdf(x)
plt.plot(x, y, c=cmap[i], ls=('-','--',':')[j], label=name)
plt.xlim(x[0], x[-1])
plt.legend(fontsize='large')
A Very Simple Example#
To get started we need something to fit. Real data is always messy and complicated. We’ll start with something very simple.
Here is a simple linear model and we’ll generate a few random points along some line. We need to identify some uncertainties but here we’ll just take them all to have the same size (here 0.1).
Plotting the data confirms that we’ve got the code working as we expect.
# a simple linear model,
def line(x, a, b):
return a + x * b
# generate random toy data with random offsets in y
np.random.seed(seed=int(time.time()))
rng = np.random.default_rng(1)
data_x = np.linspace(0, 1, 10)
data_yerr = 0.1 # could also be an array
data_y = rng.normal(line(data_x, 1, 2), data_yerr)
print(data_y)
plt.errorbar(data_x, data_y, data_yerr, fmt="ok", label="data")
Least Square Fitting#
In the example above, we are going to fit the data points to a line of the form \(\alpha + \beta x\), where \(\alpha\) and \(\beta\) are the values we would like the fit to choose. This is a choice (in this case, a very good choice). As we’ll see below, the fit parameters can be very meaningful. There is no limit on the number of fit parameters you can include. Adding more and more parameters will make the fit unstable (try it).
If you’ve run across fitting before it has probably been Least-squares fitting. This finds the best fit by minimizing this quantity:
Here:
in the case of the data we’ve generated above, \(i\) would run from 1-10
\(y_i\) are each of the individual measured values of \(y\) (so the contents of data_y in our example code),
\(\sigma_i\) are the values of the measurement uncertainties (0.1 for all \(i\) in our example code)
\(y_{\alpha,\beta}(x_i)\) are the values of the fitted function evaluated for a particular choice of \(\alpha\) and \(\beta\) at \(x_i\) values
We don’t know what values of \(\alpha\) and \(\beta\) are optimal. The least-squares fitting routine tries different fit parameters and evaluates the \(\chi^2\) value for each choice. The best choice of \(\alpha\) and \(\beta\) values correspond to the minimum value of $\chi^2(\alpha,\beta).
Looking at the expression for \(\chi^2_{\alpha,\beta}\) more closely, we see:
Minimizing the square of the deviations makes sense qualitatively because if you minimized the deviations themselves, it would be possible to get a fit that was equally off in the positive and negative direction that had a very small value of \(\chi^2\) (maybe like a horizontal line in linear example above).
The factors of \(1/\sigma_i^2\) ensure that the points with smaller uncertainty have more power in the fit. In the example above all the uncertainties are the same but that’s not true in many cases. You wouldn’t want to get a wildly different fit by adding a outlying but very imprecise point (in the example above maybe \(x=0.5\), \(y = 7 \pm 4\)).
iminuit has a inutitive interface for least squares minimization. It is based on Minuit which is widely used in high energy and nuclear physics. There are many different implementations of least-squares fitting available.
least_squares = LeastSquares(data_x, data_y, data_yerr, line)
m = Minuit(least_squares, a=0, b=0) # starting values for α and β
m.migrad() # finds minimum of least_squares function
m.hesse() # accurately computes uncertainties
So what do we see?
First the values of the parameters alpha and beta are close to the values we put in above. And the fitted curve goes through the points (mostly). This is promising and suggests we’re doing pretty good.
it also computes the Hesse errors. This is a way of computationally getting the covariance matrix.
you can find the covariance matrix right above the plot (check out week 4). The diagonal elements of the covariance matrix are the squares of the errors quoted in the table above as we expect. The matrix is symmetric (as we expect) and the off-diagonal elements should be \(\rho_{xy}\sigma_x \sigma_y\) (the second number in bold on the off-diagonal elements is just \(\rho_{xy}\)).
# draw data and fitted line
plt.errorbar(data_x, data_y, data_yerr, fmt="ok", label="data")
plt.plot(data_x, line(data_x, *m.values), label="fit")
# display legend with some fit info
fit_info = [
f"$\\chi^2$/$n_\\mathrm{{dof}}$ = {m.fval:.1f} / {m.ndof:.0f} = {m.fmin.reduced_chi2:.1f}",
]
for p, v, e in zip(m.parameters, m.values, m.errors):
fit_info.append(f"{p} = ${v:.2f} \\pm {e:.2f}$")
#you want to format this so there is generally one significant figure in the uncertainty
plt.legend(title="\n".join(fit_info), frameon=False)
plt.xlabel("x")
plt.ylabel("y");
Here we’ve got the \(χ^2/n_{dof}\), what does that mean? And where do the uncertainties come from?
In lecture 3 we had the \(\chi^2\) distribution being: $\(\Large f(x;n) = \frac{1}{{2^{n/2}\Gamma(n/2)}} x^{(n/2)-1} e^{-x/2}\)$.
Above we had: $\(\Large \chi^2 ≡ \sum_{i=1}^{n} (y_i - y(x_i))^2/\sigma_i^2\)$
It is possible to show that for \(n\) independent Gaussian random variables \(y_i\) the above summation follows the \(\chi^2\) distribution for \(n\) degrees of freedom (Cowan 10.2). For any particular dataset, we get a single value of \(\chi^2\), but if we generate new datasets independently drawn from the same Gaussian PDFs as the one we did generate, then the \(\chi^2\) values of those datasets will be drawn according to the \(\chi^2\) distribution.
The above plot has 10 data points and \(n_{dof} = 8\) in the plot. Why are these numbers not the same? Because there are 2 fit parameters (the slope and the intercept) so there are only \(10-2 = 8\) independent pieces of information in the fit.
A good fit with have \(χ^2/n_{dof} ≈ 1\). How close does it have to be? Well for a given dataset, the value of \(\chi^2\) can be calculated as above. The expectation value of the \(\chi^2\) distribution is \(n\) (recall from lecture 3) and so one should have a \(χ^2/n_{dof} ≈ 1\).
How to tell if a fit is any good#
Qualitatively#
The variance of the \(\chi^2\) distribution is \(2n\), so the relative width of the distribution \(\sigma / \mu \) gets smaller as \(n\) increases, thus \(χ^2/n_{dof}\) for a good fit should be closer to one as \(n_{dof}\) increases.
Quantitatively#
We look at the CDF of the \(\chi^2\) distribution for \(n_{dof}\) What we’re interested in is what fraction of the time would we get a larger value of \(\chi^2\) if this data are described by this parameterization?
The CDF, provides the integral of the \(\chi^2\) distribution that has a smaller \(\chi^2\) value than we got and the survival fraction is 1 - CDF for our \(\chi^2\) value.
With this now there is a probability that if these fit parameters are correct, we would have gotten a smaller \(\chi^2\) value from the data. If the CDF for our \(\chi^2\) value is very small, that means that the \(\chi^2\) value is very small and the fit parameters well describe the data. As the CDF value approaches unity, the chances that these parameter describe the data decrease (though perhaps we got unlucky).
You have to judge whether the CDF associated with a particular fit is good enough for your purposes.
In general, if the conditions above are met and many independent datasets are each fit to a correct model, the CDF values associated with the \(\chi^2\) values for each dataset should populate a uniform distribution between 0 and 1.
import scipy
from scipy.stats import chi2
pdf_demo(-0.5, 25, chi2='df=8')
print(chi2.cdf(4.0,8))
print(chi2.sf(4.0,8))
chi2_values = np.empty(1)
cdf_values = np.empty(1)
for i in range(1000):
np.random.seed(seed=int(time.time()))
rng = np.random.default_rng()
data_x = np.linspace(0, 1, 10)
data_yerr = 0.1 # could also be an array
data_y = rng.normal(line(data_x, 1, 2), data_yerr)
least_squares = LeastSquares(data_x, data_y, data_yerr, line)
m = Minuit(least_squares, a=0, b=0) # starting values for α and β
m.migrad()
m.hesse() #
if i == 1:
chi2_values[0] = m.fval
cdf_values[0] = chi2.cdf(m.fval,8)
else:
chi2_values = np.append(chi2_values,m.fval)
cdf_values = np.append(cdf_values,chi2.cdf(m.fval,8))
fig, axs = plt.subplots(1, 2)
counts, bins = np.histogram(chi2_values)
counts_cdf, bins_cdf = np.histogram(cdf_values)
axs[0].hist(bins[:-1], bins, weights=counts)
axs[1].hist(bins_cdf[:-1], bins_cdf, weights=counts_cdf)
print("mean: ", chi2_values.mean())
print("variance: ", chi2_values.var())
Reduced \(\chi^2\)#
The example above is why you may have heard that the \(\chi^2/n_{dof}\), the reduced \(\chi^2\) of a good fit should be around 1. That’s true because the mean of the \(\chi^2\) distribution is \(n\).
However, the variance of the \(\chi^2\) distribution is \(2n\) so the standard deviation of the \(\chi^2\) distribution is \(\sqrt{2n}\) so the \(\sigma / mean\) of the \(\chi^2\) distribution is \(\sqrt{2/n}\).
This means a “good” value for the \(\chi^2/n_{dof}\) depends on \(n_{dof}\) as illustrated below.
chi2_values = np.empty(1)
cdf_values = np.empty(1)
rchi2_values = np.empty(1)
npts = 100
for i in range(1000):
np.random.seed(seed=int(time.time()))
rng = np.random.default_rng()
data_x = np.linspace(0, 1, npts)
data_yerr = 0.1 # could also be an array
data_y = rng.normal(line(data_x, 1, 2), data_yerr)
least_squares = LeastSquares(data_x, data_y, data_yerr, line)
m = Minuit(least_squares, a=0, b=0) # starting values for α and β
m.migrad()
m.hesse()
ndof = npts - 2
if i == 1:
chi2_values[0] = m.fval
cdf_values[0] = chi2.cdf(m.fval,ndof)
rchi2_values[0] = m.fval/ndof
else:
chi2_values = np.append(chi2_values,m.fval)
cdf_values = np.append(cdf_values,chi2.cdf(m.fval,ndof))
rchi2_values = np.append(rchi2_values,m.fval/ndof)
fig, axs = plt.subplots(1,3)
counts, bins = np.histogram(chi2_values)
rcounts, rbins = np.histogram(rchi2_values)
counts_cdf, bins_cdf = np.histogram(cdf_values)
axs[0].hist(bins[:-1], bins, weights=counts)
axs[1].hist(rbins[:-1], rbins, weights=rcounts)
axs[2].hist(bins_cdf[:-1], bins_cdf, weights=counts_cdf)
print("mean: ", chi2_values.mean())
print("variance: ", chi2_values.var())
Plotting the Correlation Contours
The fit parameters are often correlated with each other. This can be seen in the covariance or correlation matrices but it can also be directly visualized. This can be extremely useful in seeing if a particular combination of fit parameters describes a dataset.
In 2D these are very straightforward to visualize. In higher dimensions, it is common to look at particularl slices in these contours.
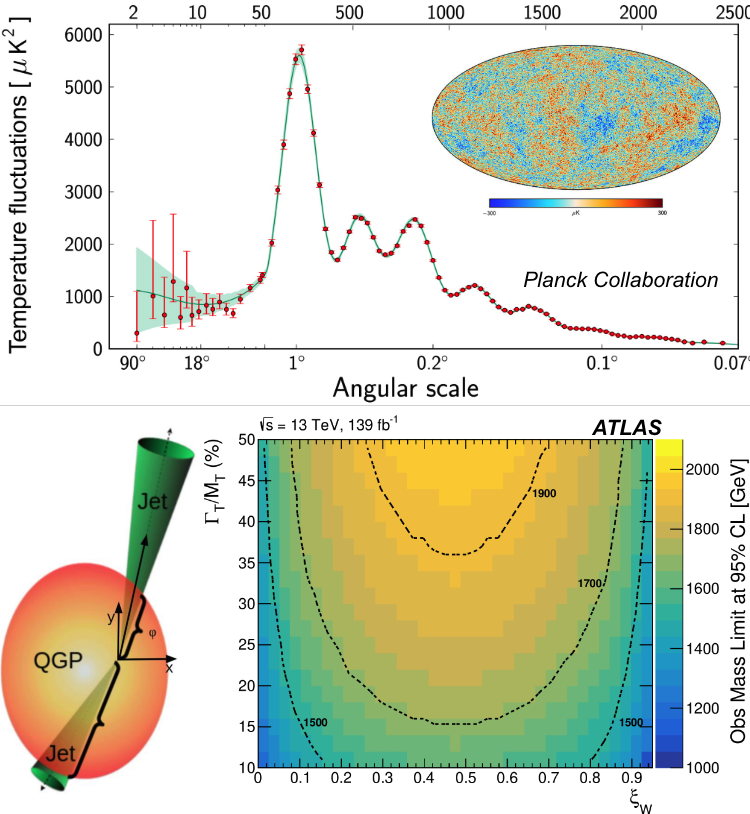
Here is an example of a contour in particle physics looking for deviations of Higgs interactions from the predictions of the Standard Model.
m.migrad()
m.draw_mncontour("a", "b")
# a simple linear model,
def line(x, a, b):
return a + x * b
def quadratic(x, a, b, c):
return a + x * b + x*x * c
least_squares = LeastSquares(data_x, data_y, data_yerr, line)
m = Minuit(least_squares, a=0, b=0) # starting values for α and β
m.migrad() # finds minimum of least_squares function
m.hesse() # accurately computes uncertainties
least_squares2 = LeastSquares(data_x, data_y, data_yerr, quadratic)
m2 = Minuit(least_squares2, a=0, b=0, c=0) # starting values for α and β, & c
m2.migrad() # finds minimum of least_squares function
m2.hesse() # accurately computes uncertainties
# generate random toy data with random offsets in y
np.random.seed(seed=int(time.time()))
rng = np.random.default_rng(1)
data_x = np.linspace(0, 1, 10)
data_yerr = 0.1 # could also be an array
data_y = rng.normal(line(data_x, 1, 2), data_yerr)
print(data_y)
plt.errorbar(data_x, data_y, data_yerr, fmt="ok", label="data")
plt.plot(data_x, line(data_x, *m.values), label="linear fit")
plt.plot(data_x, quadratic(data_x, *m2.values), label="quad fit")
ext_x = np.linspace(0, 10, 100)
plt.plot(ext_x, line(ext_x, *m.values), label="linear fit")
plt.plot(ext_x, quadratic(ext_x, *m2.values), label="quad fit")
from iminuit import cost, Minuit
import numba_stats
from numba_stats import truncnorm, truncexpon, norm, expon
import numpy as np
from matplotlib import pyplot as plt
from scipy.stats import multivariate_normal as mvnorm
# accurate numerical derivatives
%pip install jacobi
from jacobi import jacobi
A More Complicated Example#
Now let’s take a look at something more complicated. Here is a two part distribution composed of a Gaussian on an exponential distribution. Physics has many many cases like this where the peak is some kind of signal and there is a smooth is a background contribution.
xr = (0, 2) # xrange
rng = np.random.default_rng(1)
xdata = rng.normal(1, 0.1, size=1000)
ydata = rng.exponential(size=len(xdata))
xmix = np.append(xdata, ydata)
xmix = xmix[(xr[0] < xmix) & (xmix < xr[1])]
#np.histogram returns the values (n in this case) and the bin edges (xe in this case)
n, xe = np.histogram(xmix, bins=20, range=xr)
cx = 0.5 * (xe[1:] + xe[:-1])
dx = np.diff(xe)
#this plots "cx" which is the middle of every bin "xe" and n, the total counts in each bin along with the sqrt(n) as the error bar
plt.errorbar(cx, n, n ** 0.5, fmt="ok")
We’re going to take a different tactic here (though the least-squares fitting will still work with this example).
First, let’s define a PDF for our fit, \(f(y; \theta)\), \(\theta\) are the fit parameters. With our PDF and our data, we can find the parameters that maximize the probability of our data, \(y_i\), being described by the PDF we have chosen. To do that, we’ll write out the likelihood function:
We get the product here because the measurement of each \(y_i\) is independent of each other measurement, so the probabilities multiply. Often the log of \(L(\theta)\) is maximized because then the product turns into a summation.
Maximizing \(L(\theta)\) means that we want: $\(\Large \frac{\delta L}{\delta \theta_k} = 0\)$.
One (sometime) advantage here is that having a binned dataset is not required; note that below xmix which is the data itself, not a histogram, is fed into the fitter. The least squares fitting above required that the data be binned and assumes the random variables are Gaussian. For many datasets these aren’t issues, but for some datasets, especially sparse ones, binning will require either loss of information due to wide bins, or bins which have so few counts in them that the uncertainties aren’t Gaussian. In general, one wants to use binning for large datasets because using evaluating likelihoods for each measurement value will take much longer than evaluating likelihoods for each bin.
def pdf(x, z, mu, sigma, tau):
return (z * truncnorm.pdf(x, *xr, mu, sigma) +
(1 - z) * truncexpon.pdf(x, *xr, 0.0, tau))
c = cost.UnbinnedNLL(xmix, pdf)
m = Minuit(c, z=0.4, mu=1, sigma=0.2, tau=1)
m.limits["z"] = (0, 1)
m.limits["mu"] = (0, 2)
m.limits["sigma", "tau"] = (0, None)
m.migrad()
Of course we can define a binning and use that as well. (Scrolling up, see that n is a histogram.) Clearly this is an advantage as the size of the dataset grows, the number of points grows with each new measurement but the size of the histogram is fixed by the choice of binning.
def cdf(xe, z, mu, sigma, tau):
return (z * truncnorm.cdf(xe, *xr, mu, sigma) +
(1-z) * truncexpon.cdf(xe, *xr, 0, tau))
c = cost.BinnedNLL(n, xe, cdf)
m = Minuit(c, z=0.4, mu=0, sigma=0.2, tau=2)
m.limits["z"] = (0, 1)
m.limits["sigma", "tau"] = (0.01, None)
m.migrad()
We can also use the least squared fitting that we discussed above for our more complex fitting example.
def model(x, a, b, ng, d, sigma):
return a * np.exp(-x/b) + ng * np.exp(-0.5*((x-d)/sigma)**2)
c = cost.LeastSquares(cx, n, n**0.5, model)
m = Minuit(c, a=0.5, ng=1, d=1, sigma=0.1, b=1)
m.limits["sigma"] = (0.01, None)
m.migrad()